2025 MIT Health Science Forum: Treatment Effectiveness, New Biology Using AI – Enabled Nanosensor Technology (MIT Industrial Liaison Program – Startup Exchange)
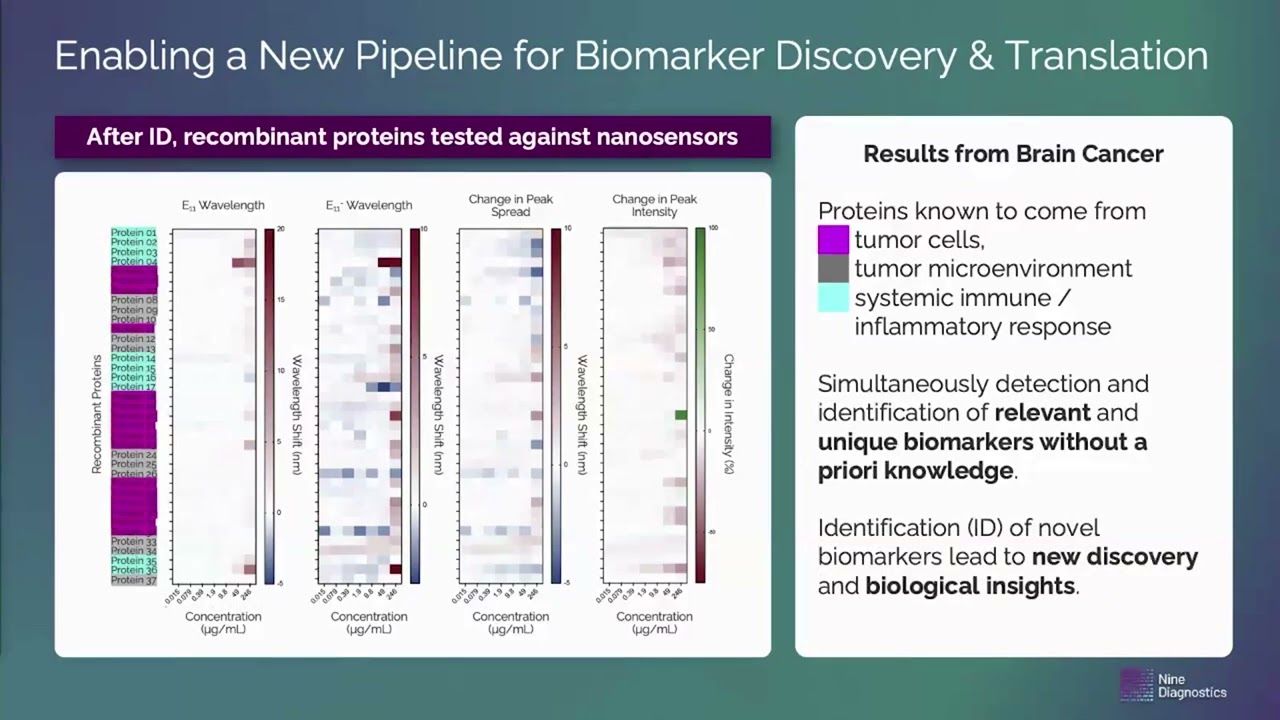
Determining Treatment Effectiveness, Discovering New Biology Using AI – Enabled Nanosensor Technology
Freddy Nguyen
CEO & Co-Founder, Nine Diagnostics
See more of the 2025 MIT Health Science Forum here: https://ilp.mit.edu/Health25
Cancer

Cancer research increasingly relies on optical and molecular technologies to enable earlier detection, more precise surgical intervention, and real-time monitoring of therapeutic response. Techniques such as fluorescence spectroscopy, Raman scattering, and optical coherence tomography (OCT) provide noninvasive access to structural, biochemical, and functional information at cellular and molecular scales. Within this space, research efforts have focused on integrating these modalities into translational platforms for oncology. Contributions include the development of spectroscopic systems for detecting epithelial precancers, intraoperative OCT technologies for margin and lymph node assessment, and multimodal contrast agents for molecular imaging. More recently, implantable carbon nanotube–based nanosensors have been used to monitor chemotherapeutic delivery and tumor microenvironment dynamics in vivo. These multidisciplinary innovations span breast, brain, pancreatic, cervical, oral, and gastrointestinal cancers, with broad applicability across both solid and hematologic malignancies.
Transfusion Medicine

Transfusion medicine is a clinical and laboratory discipline focused on the safe administration of blood and its components, encompassing immunohematology, product selection, transfusion reactions, and data-driven stewardship. Its role expanded during the COVID-19 pandemic with the emergence of convalescent plasma (CCP) as a potential therapy. Investigations characterized neutralizing antibody responses in donor plasma and evaluated CCP efficacy through propensity-matched trials in severe COVID-19. Additional research quantified adverse event rates following CCP transfusion, informing risk-benefit frameworks and institutional practice. Studies also examined pediatric O-negative red cell utilization across hospitals, revealing variation and opportunities for improved allocation. These contributions—conducted in collaboration with Mount Sinai and the New York State Department of Health—highlight the integration of immunologic profiling, transfusion safety surveillance, and clinical informatics to optimize transfusion practices and support pandemic response.
O blood usage trends in the pediatric population 2015–2019 A multi-institutional analysis

Background
In 2019, AABB released the bulletin “Recommendations on the Use of Group O Red Blood Cells” in which the recommendations about pediatric and neonatal blood transfusions were limited. Eight U.S. pediatric hospitals sought to determine trends in pediatric group O blood use and clarify which pediatric populations receive group O blood transfusions despite a non-group O blood type.
Study Design and Methods
Eight U.S.-based institutions serving a pediatric population provided data from their respective Electronic Health Records. Data submitted included unit blood type, patient blood type, patient age, sex, and discharge diagnosis. If the discharge diagnosis was not available, the admitting diagnosis was substituted. GPT-4 was used to sort diagnoses into categories for analysis. Data were visualized using a series of alluvial plots, spaghetti plots, and tables. Tables were stratified on variables of interest (blood type, age, sex, diagnosis) to explore O blood type distribution among different patient populations.
Results
A total of 142,227 discrete transfusion events were identified, of which 52,731 recipients were non-O blood type. Overall, 35,575 transfusion events of O blood went to A, B, or AB blood type recipients (67%). Additionally, 26% of Rh(D) negative transfusion events went to recipients who were Rh(D) positive. Top diagnostic categories for receiving O blood type were cardiovascular disorders (22%) and sickle cell anemia (15%).
Discussion
This study highlights opportunities to address O blood supply challenges by identifying where non-O blood may be utilized safely in the vulnerable pediatric population.
Nine Diagnostics Joins American Cancer Society’s BrightEdge Entrepreneurs Program

Nine Diagnostics, a leader in AI-enabled nanosensor technology, has been selected to participate in the American Cancer Society’s BrightEdge Entrepreneurs Program, a highly selective initiative designed to accelerate the most promising oncology-focused startups. This selection marks another significant milestone for Nine Diagnostics as it continues to drive innovation in cancer treatment selection, dosing, optimization, and monitoring.
Nine Diagnostics Selected for Merck Digital Sciences Studio Cohort 3

Arnold and Mabel Beckman Foundation – March 30, 2017
2017 Beckman Postdoctoral Fellow
Massachusetts Institute of Technology
Research: Development of nanosensors for in-vivo monitoring of cancer therapeutics
Freddy Nguyen, MD/PhD and Nine Diagnostics Win Novo Nordisk Golden Ticket at Pitch Event

Arnold and Mabel Beckman Foundation – March 30, 2017
2017 Beckman Postdoctoral Fellow
Massachusetts Institute of Technology
Research: Development of nanosensors for in-vivo monitoring of cancer therapeutics
Nine Diagnostics Wins Novo Nordisk Golden Ticket for LabCentral Residency

Arnold and Mabel Beckman Foundation – March 30, 2017
2017 Beckman Postdoctoral Fellow
Massachusetts Institute of Technology
Research: Development of nanosensors for in-vivo monitoring of cancer therapeutics
Fluorescence-based detection of protein aggregation and fiber optic-based benchtop instrument

Laser Fluence Dependent Modulation of Single Walled Carbon Nanotube Photoluminescence

There is a pressing need for sensors and assays to monitor chemotherapeutic activity within the human body in real time to optimize drug dosimetry parameters such as timing, quantity, and frequency in an effort to maximize efficacy while minimizing deleterious cytotoxicity. Herein, we develop near-infrared fluorescent nanosensors based on single walled carbon nanotubes for the chemotherapeutic Temozolomide (TMZ) and its metabolite 5-aminoimidazole-4-carboxamide using Corona Phase Molecular Recognition as a synthetic molecular recognition technique. The resulting nanoparticle sensors are able to monitor drug activity in real-time even under in vivo conditions. Sensors can be engineered to be biocompatible by encapsulation in poly(ethylene glycol) diacrylate hydrogels. Selective detection of TMZ was demonstrated using U-87 MG human glioblastoma cells and SKH-1E mice with detection limits below 30 μM. As sensor implants, we show that such systems can provide spatiotemporal therapeutic information in vivo, as a valuable tool for pharmacokinetic evaluation. Sensor implants are also evaluated using intact porcine brain tissue implanted 2.1 cm below the cranium and monitored using a recently developed Wavelength-Induced Frequency Filtering technique. Additionally, we show that by taking the measurement of spatial and temporal analyte concentrations within each hydrogel implant, the direction of therapeutic flux can be resolved. In all, these types of sensors enable the real time detection of chemotherapeutic concentration, flux, directional transport, and metabolic activity, providing crucial information regarding therapeutic effectiveness.
Molecular Recognition and In Vivo Detection of Temozolomide and 5-Aminoimidazole-4-carboxamide for Glioblastoma Using Near-Infrared Fluorescent Carbon Nanotube Sensors

There is a pressing need for sensors and assays to monitor chemotherapeutic activity within the human body in real time to optimize drug dosimetry parameters such as timing, quantity, and frequency in an effort to maximize efficacy while minimizing deleterious cytotoxicity. Herein, we develop near-infrared fluorescent nanosensors based on single walled carbon nanotubes for the chemotherapeutic Temozolomide (TMZ) and its metabolite 5-aminoimidazole-4-carboxamide using Corona Phase Molecular Recognition as a synthetic molecular recognition technique. The resulting nanoparticle sensors are able to monitor drug activity in real-time even under in vivo conditions. Sensors can be engineered to be biocompatible by encapsulation in poly(ethylene glycol) diacrylate hydrogels. Selective detection of TMZ was demonstrated using U-87 MG human glioblastoma cells and SKH-1E mice with detection limits below 30 μM. As sensor implants, we show that such systems can provide spatiotemporal therapeutic information in vivo, as a valuable tool for pharmacokinetic evaluation. Sensor implants are also evaluated using intact porcine brain tissue implanted 2.1 cm below the cranium and monitored using a recently developed Wavelength-Induced Frequency Filtering technique. Additionally, we show that by taking the measurement of spatial and temporal analyte concentrations within each hydrogel implant, the direction of therapeutic flux can be resolved. In all, these types of sensors enable the real time detection of chemotherapeutic concentration, flux, directional transport, and metabolic activity, providing crucial information regarding therapeutic effectiveness.
An Accessible, Efficient, and Accurate Natural Language Processing Method for Extracting Diagnostic Data from Pathology Reports

Analysis of diagnostic information in pathology reports for the purposes of clinical or translational research and quality assessment/control often requires manual data extraction, which can be laborious, time-consuming, and subject to mistakes.
Objective
We sought to develop, employ, and evaluate a simple, dictionary- and rule-based natural language processing (NLP) algorithm for generating searchable information on various types of parameters from diverse surgical pathology reports.
Design
Data were exported from the pathology laboratory information system (LIS) into extensible markup language (XML) documents, which were parsed by NLP-based Python code into desired data points and delivered to Excel spreadsheets. Accuracy and efficiency were compared to a manual data extraction method with concordance measured by Cohen’s κ coefficient and corresponding P values.
Results
The automated method was highly concordant (90-100%, P<.001) with excellent inter-observer reliability (Cohen’s κ: 0.86-1.0) compared to the manual method in 3 clinicopathologic research scenarios, including squamous dysplasia presence and grade in anal biopsies, epithelial dysplasia grade and location in colonoscopic surveillance biopsies, and adenocarcinoma grade and amount in prostate core biopsies. Significantly, the automated method was 24-39 times faster and inherently contained links for each diagnosis to additional variables such as patient age, location, etc., which would require additional manual processing time.
Conclusions
A simple, flexible, and scaleable NLP-based platform can be used to correctly, safely, and quickly extract and deliver linked data from pathology reports into searchable spreadsheets for clinical and research purposes.
